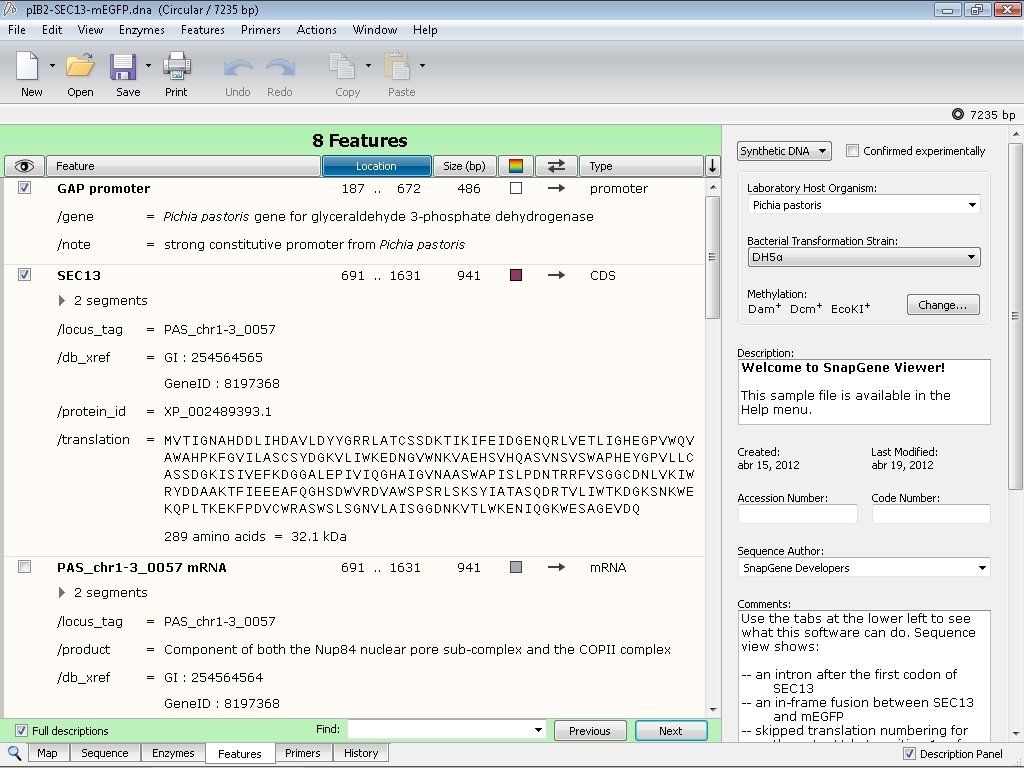
The yellow highlighted area contains the DNA fragment cleaved out following NdeI/XhoI digestion of the pET26b plasmid. PET26b plasmid previously digested with NdeI and XhoI =? To answer this question let us zoom in on the pET26b plasmid map and look at the actual sequence surrounding the NdeI and XhoI restriction enzyme DNA recognition sites: lacI – a gene that expresses the Lac repressor, a protein used to control the T7/ lac promoter.Īnd so, based on this map we can say the following:.Ori – the origin of replication for the pET26b plasmid.always on) and is used as a selectable marker for bacterial transformations. kanR gene, selectable marker – the kanamycin resistance gene is situated downstream of a constitutive promoter (i.e.Why? Because this His 6x-tag will be used later for protein purification. His 6x-tag – this is an engineered tag consisting of the codon for the amino acid histidine repeated six times.This indicates the bp position of the cut site for each RE. Note the bp position in brackets beside each RE. NdeI and XhoI – the two Restriction Enzymes (RE) we are using to clone our folA gene downstream of the T7/ lac promoter.We are cloning our folA gene downstream of this promoter. we control when this promoter is off or on). the T7/ lac promoter – this is the main event! An inducible promoter (i.e.The pET26b plasmid is a circular, covalently-closed double-stranded DNA, 5360 bp in length. Note the different features of the plasmid and their base pair (bp) location on the plasmid. This is a map generated from the actual pET26b DNA sequence. This is a paid software but it does have a basic, freeware version called SnapGene viewer. There are many molecular cloning software programs available, and some are free to use. 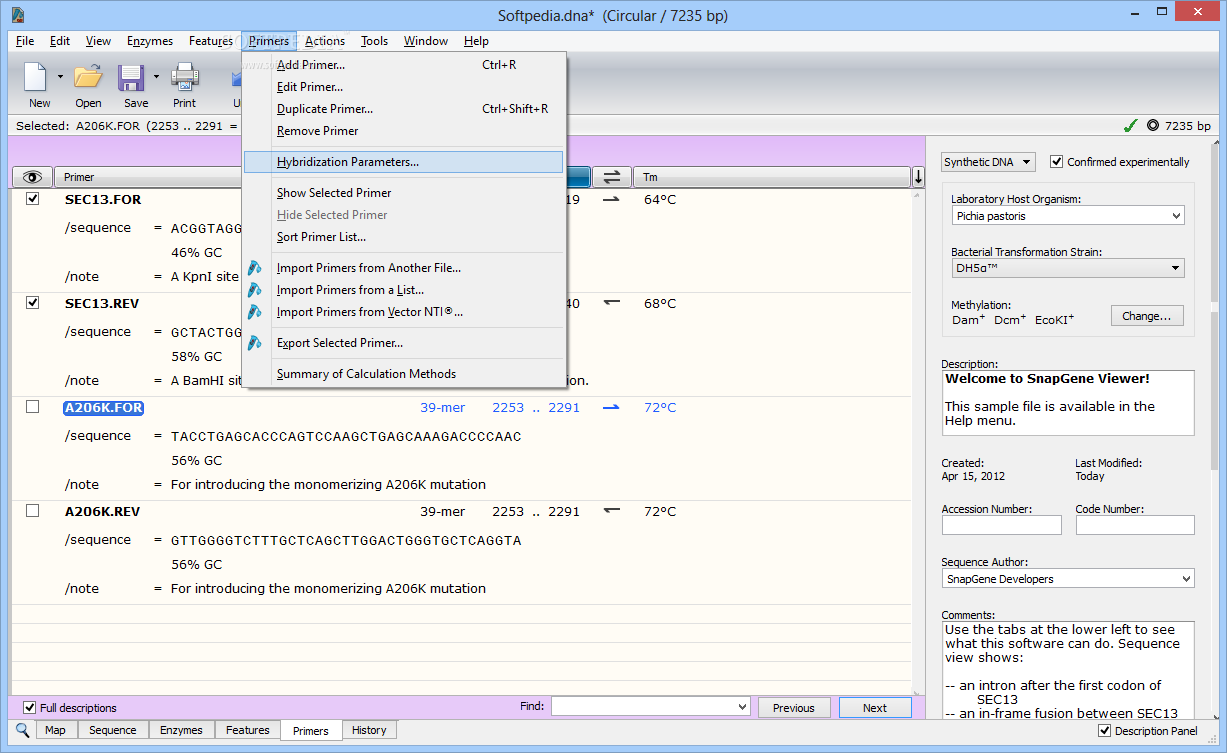
The sequence can be processed using molecular cloning software. One such location is the AddGene portal: pET26b sequence The pET26b plasmid sequence can be sourced from many locations. OK, let’s unpack this, starting with the pET26b plasmid. coli folA gene (and sequence of the DNA primers used in the PCR). This question is actually quite complex and requires knowledge of the DNA sequence for both pET26b plasmid and the E. If you wish to learn more about our Team Think Tanks, click here. Please check out Team Think 3: Restriction Enzyme Digests located in the Teacher Resources section. Helping students understand how to determine DNA fragment sizes following RE digestion reactions makes for a great Team Think Tank. Chapter 2 background – expected DNA band sizes
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
